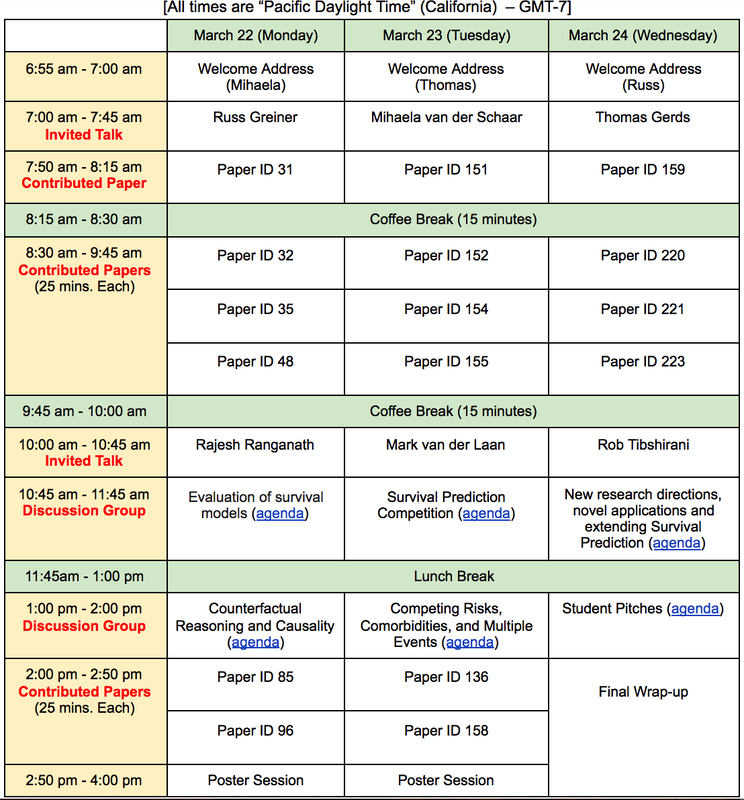
For details of each paper, see the proceedings of this symposium at MLR press [http://proceedings.mlr.press/v146/].
Invited Talks

Speaker: Prof. Russ Greiner, University of Alberta
Title: An Effective Way to Estimate an Individual's Survival Distribution
Abstract: An accurate estimate of a patient’s survival time can help determine the appropriate treatment and care of that patient. Some common approaches to survival analysis estimate a patient’s risk scores; others estimate a patient’s 5-year survival probability, or a population’s survival distribution; however, none of these provides a way to estimate an individual’s expected survival time. This motivates an alternative class of tools that can learn models that estimate a subject’s survival probability at each time -- ie, an individual survival distribution (ISD) -- from which one can then estimate that subject’s expected survival time. After describing such ISD models and explaining how they differ from standard models, this presentation then discusses standard ways to evaluate such models, then motivates and defines a novel approach, 'D-Calibration', which determines whether a model's probability estimates are meaningful. We also discuss how these measures differ, and use them to evaluate several ISD prediction tools over a range of real-world survival data sets -- demonstrating, in particular, that one tool, MTLR, provides survival estimates that are helpful for patients, clinicians and researchers.
Title: An Effective Way to Estimate an Individual's Survival Distribution
Abstract: An accurate estimate of a patient’s survival time can help determine the appropriate treatment and care of that patient. Some common approaches to survival analysis estimate a patient’s risk scores; others estimate a patient’s 5-year survival probability, or a population’s survival distribution; however, none of these provides a way to estimate an individual’s expected survival time. This motivates an alternative class of tools that can learn models that estimate a subject’s survival probability at each time -- ie, an individual survival distribution (ISD) -- from which one can then estimate that subject’s expected survival time. After describing such ISD models and explaining how they differ from standard models, this presentation then discusses standard ways to evaluate such models, then motivates and defines a novel approach, 'D-Calibration', which determines whether a model's probability estimates are meaningful. We also discuss how these measures differ, and use them to evaluate several ISD prediction tools over a range of real-world survival data sets -- demonstrating, in particular, that one tool, MTLR, provides survival estimates that are helpful for patients, clinicians and researchers.

Speaker: Prof. Thomas A. Gerds, University of Copenhagen
Title: Medical risk prediction models: Can't the computer take care of all this?
Abstract: The computer cannot detect the target of prediction and deal with the
obstacles of the data: No, it is not OK to delete all the weird cases such as death, or loss to followup, or missing predictor value. But, even if we communicate the full medical story correctly, current machine learning methods tend to produce biologically implausible models that even depend on the random seed. You might think that there is one correct way to build a risk prediction model. This is not true, but different strategies have their strengths and weaknesses. We have weak learners, strong learners and stacking (AKA super learner). We have fully specified, i.e., expert selected, semi-parametric models and penalized regressions which have hyperparameters too. In this talk, I describe a general framework for the comparison of rival modelling strategies with respect to their performance of predicting the risks that a medical event occurs within t-years in the presence of competing risks and based on right-censored data.
Title: Medical risk prediction models: Can't the computer take care of all this?
Abstract: The computer cannot detect the target of prediction and deal with the
obstacles of the data: No, it is not OK to delete all the weird cases such as death, or loss to followup, or missing predictor value. But, even if we communicate the full medical story correctly, current machine learning methods tend to produce biologically implausible models that even depend on the random seed. You might think that there is one correct way to build a risk prediction model. This is not true, but different strategies have their strengths and weaknesses. We have weak learners, strong learners and stacking (AKA super learner). We have fully specified, i.e., expert selected, semi-parametric models and penalized regressions which have hyperparameters too. In this talk, I describe a general framework for the comparison of rival modelling strategies with respect to their performance of predicting the risks that a medical event occurs within t-years in the presence of competing risks and based on right-censored data.

Speaker: Prof. Mihaela van der Schaar, University of Cambridge, UCLA, Turing Institute
Title: Survival Analysis in the Era of Machine Learning: Is there life after Cox?
Abstract: For several decades, Cox Regression has represented the gold standard for survival analysis. My talk will focus on several areas where our recent work on machine learning has led to significant improvements over Cox Regression: 1) Automated machine learning, 2) Competing risks, 3) Time-series data, and 4) Transfer learning. I will also discuss the application of these methods in a variety of healthcare scenarios. These results suggest that machine learning approaches might replace Cox regression as the gold standard for some kinds of problems.
Title: Survival Analysis in the Era of Machine Learning: Is there life after Cox?
Abstract: For several decades, Cox Regression has represented the gold standard for survival analysis. My talk will focus on several areas where our recent work on machine learning has led to significant improvements over Cox Regression: 1) Automated machine learning, 2) Competing risks, 3) Time-series data, and 4) Transfer learning. I will also discuss the application of these methods in a variety of healthcare scenarios. These results suggest that machine learning approaches might replace Cox regression as the gold standard for some kinds of problems.

Speaker: Prof. Rajesh Ranganath, New York University
Title: Deep Survival Analysis
Abstract: Survival models that predict the time to an event are ubiquitous in medicine. In the last several years there has been a large body of work developing deep learning methods for survival analysis. Survival analysis, however, can be tricky. Unlike many traditional prediction problems in deep learning, the target being predicted in survival analysis, the time to an event, can be hidden. In this talk, I will lay out challenges that arise with deep survival models in practice such as calibration, missing data, and interpretability along with some of our solutions for them.
Title: Deep Survival Analysis
Abstract: Survival models that predict the time to an event are ubiquitous in medicine. In the last several years there has been a large body of work developing deep learning methods for survival analysis. Survival analysis, however, can be tricky. Unlike many traditional prediction problems in deep learning, the target being predicted in survival analysis, the time to an event, can be hidden. In this talk, I will lay out challenges that arise with deep survival models in practice such as calibration, missing data, and interpretability along with some of our solutions for them.

Speaker: Prof. Mark van der Laan, University of California, Berkeley
Title: Targeted Maximum Likelihood Estimation of the Causal Effect of Treatment on Survival
Abstract: In this talk we provide an overview of TMLE, and the highly adaptive lasso (HAL), where the latter machine learning algorithm guarantees the asymptotic efficiency of the TMLE. In particular, we present the universal least favorable path along which the TMLE update is achieved in one single most efficient step, resulting in the one-step TMLE. This also allows the TMLE to target infinite dimensional target parameters such as treatment specific survival functions, thereby guaranteeing a compatible substiution estimator of such functions respecting the monotonicity constraints. We discuss the resulting causal inference methods for survival data based on discrete and continuous time to event data. (Joint work with Helene Rijtgaard, Thomas Gerds, Susan Gruber)
Title: Targeted Maximum Likelihood Estimation of the Causal Effect of Treatment on Survival
Abstract: In this talk we provide an overview of TMLE, and the highly adaptive lasso (HAL), where the latter machine learning algorithm guarantees the asymptotic efficiency of the TMLE. In particular, we present the universal least favorable path along which the TMLE update is achieved in one single most efficient step, resulting in the one-step TMLE. This also allows the TMLE to target infinite dimensional target parameters such as treatment specific survival functions, thereby guaranteeing a compatible substiution estimator of such functions respecting the monotonicity constraints. We discuss the resulting causal inference methods for survival data based on discrete and continuous time to event data. (Joint work with Helene Rijtgaard, Thomas Gerds, Susan Gruber)

Speaker: Prof. Rob Tibshirani and Erin Craig, Stanford University
Title: Survival Analysis as a Classification Problem
Abstract: We present a method for treating survival analysis as a classification problem. This method uses a “stacking” idea that collects features and outcomes of the survival data in a large data frame, and treats it as a classification problem. In this framework, various statistical learning algorithms (including logistic regression, random forests, gradient boosting machines and neural networks) can be applied to estimate parameters and make predictions.
Title: Survival Analysis as a Classification Problem
Abstract: We present a method for treating survival analysis as a classification problem. This method uses a “stacking” idea that collects features and outcomes of the survival data in a large data frame, and treats it as a classification problem. In this framework, various statistical learning algorithms (including logistic regression, random forests, gradient boosting machines and neural networks) can be applied to estimate parameters and make predictions.